Note: I’m assuming if you are reading this post that you’ve first read this post, summarizing the concept. The below is a companion piece with some expanded details about the concepts and more examples of how I think addressing this bottleneck will help make a difference.
—
You might hold the perspective that in the growing era of AI, there’s too much noise already. Slopocalypse. (Side note: this entire post, and my other post, was written by me, a human.) That’s not the biggest problem in healthcare, though: in both research and clinical care, so much critical data is simply not collected. We are missing sooooo much important data. Some of this is an artifact of past clinical trial design and how it was hard to collect or analyze or store the data; there were less established norms around data re-use; and some of it is a “collect the bare minimum to study the endpoints because that’s all we are supposed to do” phenomenon.
Nowadays, we should be thinking about how to make data and insights from clinical work and research available to AI, too, because AI will increasingly be used by humans to sort through what is known and where the opportunities are, plus make cross-domain connections that humans have been missing. And we need to be thinking about whether the incentives are set up correctly (spoiler: they’re not) to make sure all available data is able to be collected and shared or at least stored for humans and AI to have access to for future insight. What is studied is also ‘what is able to be funded’, which is disproportionately things that come to the commercial market eventually after regulatory approval.)
“But what’s the point?” you ask. “If no one is looking at the data from this study, why collect it?” Because the way we design studies – to answer a very limited-scope question, i.e. safety and efficacy for labeling claims and regulatory approval – is very different than the studies we need around optimizing and personalizing treatments. If you look at really large populations of people accessing a treatment – think GLP-1 RA injectables, for example – you eventually start to see follow-on studies around optimization and different titration recommendations. But most diseases and most treatments aren’t for a population even 1/10th the size of people accessing those meds. So those studies don’t get done. Doctors don’t necessarily pay attention to this; patients may or may not report relevant data about this back to the doctor (and again even if they do: doctors don’t mentally or physically store this data, often); and the signal is missed because we don’t capture this ‘noise’.
This means people with rare diseases, undiagnosed diseases, atypical or seronegative diseases, unusual responses to treatments, multiple conditions (comorbidities), or any other scenario that results in them being part of a small population and where customizing and individualizing their care is VERY important…they have no evidence-based guidance. Clinicians do the best they can to interpret in the absence of evidence or guidelines, but no wonder patients often turn to LLMs (AI) for additional input and contextualization and discussion of the tradeoffs and pros and cons of different approaches.
We have to: the best available human (our clinicians) may not have the relevant expertise; or they may not be available at all; or they may be biased (even subconsciously) or forgetful of critical details or not up to date on the latest evidence; or they may be faced with a truly novel situation and they don’t have the skills to address it because they’re used to cookie-cutter standardized cases. This is not a dig at clinicians, but I recognize that we now have tools that can address some of the existing flaws in our human-based healthcare system. I keep talking about how we need to recognize that evaluations of AI in healthcare shouldn’t treat the status quo as the baseline to defend, because the status quo itself has problems, as I just described.
And AI has some of the same problems. (Except for the ‘not available’ part, unless you consider the lower-utility free access models to mean that the more advanced, thinking-based models are ‘not available’ because of cost access barriers). An AI may not have any training data on a rare disease… because nothing exists. It may drop information out of the context window, and we may not realize that this has happened (i.e., it ‘forgets’ something). Usually these are critiques of AI, juxtaposed against the implication that humans are better. But notice that these critiques are the same of humans, too! This happens all the time with human clinicians in healthcare. A human can’t make decisions on data that doesn’t exist in the world, either!
So…how do we “fix” AI? Or, how do we fix human healthcare? We should be asking BOTH questions. Maybe the answer is the same: increase the noise so we increase the signal. Argue about the ratio later (of signal:noise), but increase the amount of everything first. That is most important.
How might we do this? I have been thinking about this a lot, and Astera recently posted an essay contest asking for ideas that don’t fit the current infrastructure. (That’s why I’m finally writing this up, not because I think I’ll win the essay contest, but mostly because i’s an opportunity for people to consider whether/not this type of solution is a good idea as a separate analysis from “well, who is going to fund THAT?”. Let’s discuss and evaluate the idea, or riff on it, without being bogged down by the ‘how exactly it gets funded and managed over time’.)
I think we should create some kind of digitally-managed platform/ecosystem to do the following:
- Incentivize and facilitate written (AI-assisted allowed) output of everything. Case reports and scenarios of people and what they’re facing. All the background information that might have contributed. All the data we have access to that is related to the case, plus other passive, easy-to-collect data (that may already be collected, e.g., wearables and phone accelerometer data) even if it does not appear to be related. Patient-perspective narratives, clinical interpretations, clinical data, wearable data – all of this.
A.) And, be willing to take variable types of data even from the same study population. For example, as a person with type 1 diabetes, I have 10+ years of CGM data. For a future study on exocrine pancreatic insufficiency, for example, not everyone will have CGM data, but because CGM data is increasingly common in people with T2D (a much larger population) and in the general population, a fraction of people will have CGM data and a fraction of people are willing to share it. We should enable this, even if it’s not for a primary endpoint analysis on the EPI study and even if only a fraction of people choose to share it – it might still be useful for subgroup analysis, determining power for a future study where it is part of the protocol, and identifying new research directions!
- Host storage somewhere. There can be some automated checking against identifiable information and sanity checking what’s submitted into the repository, plus consent (either self-consent or signed consent forms for this purpose.)
- Provide ‘rewards’ as incentives for inputting cases and data.
A) Rewards might differ for patients self-submitting data and researchers and clinicians inputting data and cases. Maybe it’s one type of reward for patients and a batch reward of a free consult or service of some kind for researchers/clinicians (e.g. every 3 case reports submitted = 1 credit, credits can be used toward a new AI tool or token budget for LLMs or human expertise consult around study design, or all kinds of things.)
B) Host data challenges where different types of funders incentivize different disease groups or families of conditions to be added, to round out what data is available in different areas.
- Enable AI and human access (including to citizen scientists/independent researchers who don’t have institutions) to these datasets, after self-credentialing and providing a documented use case for why the data is being accessed.
A) Require anyone accessing the dataset to make their analysis code open source, or otherwise openly available for others to use, and to make the results available as well.
B) Tag anytime an individual dataset is used by a project, that way individuals with self-donated data (or clinicians submitting a case) can revisit and see if there are any research insights or data analysis that applies to their case.
C) Provide starter projects to show how the data can be used for novel insight generation. For example, develop sandboxes for different types of datasets with existing ‘lab notebooks’, so to speak, to onboard people to different datasets/groups of data, different types of analyses they might do, and to cut down on the environment setup barriers to getting started to analyzing this data.
D) Facilitate outreach to institutions, disease-area patient nonprofits and advocacy groups both to solicit data inputs AND to use the data for generating outputs.
Why should we do this?
- Clinical trials do not gather all of the possible useful data, because it’s not for their primary/secondary endpoints. Plus, they would have to clean it upon collection. Plus storage cost. Plus management decisions. Etc. There’s a lot of real barriers there, but the result is that clinical trials do not capture enough or the most possible relevant data. So we need a different path.
- Clinicians/clinical pathways don’t have ways to review data, so they avoid doing it. There’s no path to “submit this data to my record but I don’t expect you to look at it” in medical charts (but there should be). So we need a different path.
- Disease groups/organizations sometimes host and capture datasets, but like clinical trials, this is limited data; it often comes through clinical pathways (further limiting participation and diversity of the data); it doesn’t include all the additional data that might be useful. So we need a different path.
- Some patients do publish or co-author in traditional journals, but it’s a limited, self-selected group. Then there are the usual journal hurdles that filter this down further. Not everyone knows about pre-prints. And, not everyone knows how useful their data can be – either as an n=1 alone or as an n=1*many for it to be worth sharing. A lot of people’s data may be in video/social media formats inaccessible to LLMs (right now) or locked in private Facebook groups or other proprietary platforms. So we need a different path.
Thus, the answer to ‘why we should do this’ is recognizing that: AI does not create magic out of nowhere. It relies on training data and extrapolating from that, plus web search. If it’s not searchable or findable, and it’s not in the training data, it’s not there. Yes, even the newer models and prompting it to extrapolate doesn’t solve all of these problems. What AI can do is shaped disproportionately by formal literature, institutional documents, and whatever scraps of publicly available content remain. If we want to shape what’s possible in the future, we need to start now by shaping and collecting the inputs to make it happen. We can’t fix the past trial designs but we can start to fill in the gaps in the data!
Here is an example of how this might work and why it matters to build and incentivize this type of data sharing.
- If we build and incentivize data sharing, the following individuals might self-donate their data and write-ups, or a clinician might submit them. Later, someone might analyze the data and put the pieces together.
- Someone with type 1 diabetes (an autoimmune condition) and this new-onset muscle-related issue. Glucose levels are well-managed, and they don’t have neuropathy/sensory issues (which are common complications of decades of living with type 1 diabetes), and muscle damage and inflammation markers are normal. MRIs are normal. The person submits their data which includes their lab tests, clinical chart notes (scrubbed for anonymity), a patient write-up of what their symptoms are like, and exported data from their phone with years of motion/activity data.
- A different person with Sjogren’s disease (also an autoimmune condition) and a similar new-onset muscle-related issue. There are known neurological manifestations or associations with Sjogren’s, but more typically those are small-fiber neuropathy or similar. The symptoms here are not sensory or neuropathy. MRIs don’t show inflammation or muscle atrophy. Their clinician is stumped, but knows they don’t see a lot of patients with Sjogren’s and wonders if there is a cohort of people with Sjogren’s facing this. The clinician asks the patient and gains consent; scrubs the chart note of identifying details, and submits the chart notes, labs, and a clinician summary describing the situation.
- Later, a researcher (either a traditional institutional-affiliated researcher OR a citizen scientist, such as a third person with a new-onset muscle-related issue) decides to investigate a new-onset muscle-related issue. They register their hypothesis: that there may be a novel autoimmune condition that results in this unique muscle-related issue as a neuromuscular disease (that’s not myasthenia gravis or another common NMJ disease) that shows up in people with polyautoimmunity (multiple autoimmune conditions), but we don’t know which antibodies are likely correlated with it. The research question is to find people in the platform with similar muscle-related conditions and explore the available lab data to help find what might be the connecting situation and classify this disease or better understand the mechanism.
- They are granted access to the platform, start analyzing, and come across an interesting correlation with antibody X, which is considered a standard autoimmune marker but one that doesn’t differentiate by disease, and which does highly correlate with this possibly novel muscle condition when it exists and is elevated in people with at least one existing autoimmune condition and these muscle-symptoms that are seronegative for every other autoimmune and neurologic condition. Further, by reading the clinical summary related to patient B and the self-written narrative from patient A, it becomes clear that this is likely neuromuscular – stemming from transmission failure along the nerves – and that the muscles are the ‘symptom’ but not the root cause of disease. This provides an avenue for a future research protocol to 1) allow these types of patients to be characterized into a cohort so this can be determined whether it is a novel disease or a subgroup of an adjacent condition (e.g., a seronegative subgroup); 2) track whether the antibody levels are treatment-sensitive or not or stay elevated always; and 3) cohorts of treatments that can be trialed off-label because they work in similar NMJ diseases even though the mechanism isn’t identical.
- The mechanism for this novel disease isn’t proven, yet, but in the face of all the previous negative lab data and neurological testing patient A and patient B have experienced, it narrows it down from “muscle or neuromuscular” which is a significant improvement from their previous situations. Plus, this provides pathways for additional characterization; research; and eventual treatment options to explore versus the current dead ends both patients (and their clinical teams) are stuck in. And because there are no good clinical pathways for these types of undiagnosed cases, this type of insight development across multiple cases would not have occurred at all without this database of existing data.
2. The above is a small-n example, but consider a large dataset where there are hundreds or thousands of people with CGM data submitted. Plus meal tracking data, because people can export and provide that from whatever meal-logging apps some of them happen to use.
- By analyzing this big dataset, an individual researcher could hypothesize that they could build a predictor to identity the onset of exocrine pancreatic insufficiency, which can occur in up to 10% of the general population (more frequently in older adults, people with any type of diabetes, as well as other pancreas-related conditions like pancreatitis, different types of cancer, etc), by comparing increases in glucose variability that correlate with a change in dietary consumption patterns, notably around decreasing meal size and eventually lowering the quantity of fat/protein consumed. (These are natural shifts people make when they notice they don’t feel good because their body is not effectively digesting what they are eating). They analyze and exclude the effect of GLP1-RA’s and other medications in this class: the effect size persists outside of medication usage patterns. This can later be validated and tested in a prospective clinical trial, but this dataset can be used to identify what level of correlation between meal consumption change and glucose variability change happen over what period of time in order to power a high-quality subsequent clinical trial. This may lead to an eventual non-invasive method to diagnose exocrine pancreatic insufficiency through wearable and meal-tracking data. (None exists today: only a messy stool test that no one wants to do and is hampered by other issues for accuracy.)
–
These are two examples and show how even small-n data or a large dataset where there are additional subgroups with additional datasets can be useful. In the past, it’s been a “we need X, Y, Z data from everyone. A, B, C would be nice to have, but it’s hard and not everyone will share it (or is willing to collect it due to burden), so we won’t enable it to be shared for the 30% of people who are willing”. Thus, we lose the gift and contributions from the people who are able and willing to share that data. Sometimes that 30% is a small n but that small n is >0 and may be the ONLY data that will eventually answer an important future research question.
 We are missing so much, because we don’t collect it. So, we should collect it. We need a platform to do this outside of any single disease group patient registry; we need to support clinician and patient entries into this platform; we need to support intake of a variety of types of data; and we need to have low (but sensible) barriers to access so individuals (citizen scientists, patients themselves) can leverage this data alongside traditional researchers. We need all hands on deck, and we need more data collected.
We are missing so much, because we don’t collect it. So, we should collect it. We need a platform to do this outside of any single disease group patient registry; we need to support clinician and patient entries into this platform; we need to support intake of a variety of types of data; and we need to have low (but sensible) barriers to access so individuals (citizen scientists, patients themselves) can leverage this data alongside traditional researchers. We need all hands on deck, and we need more data collected.

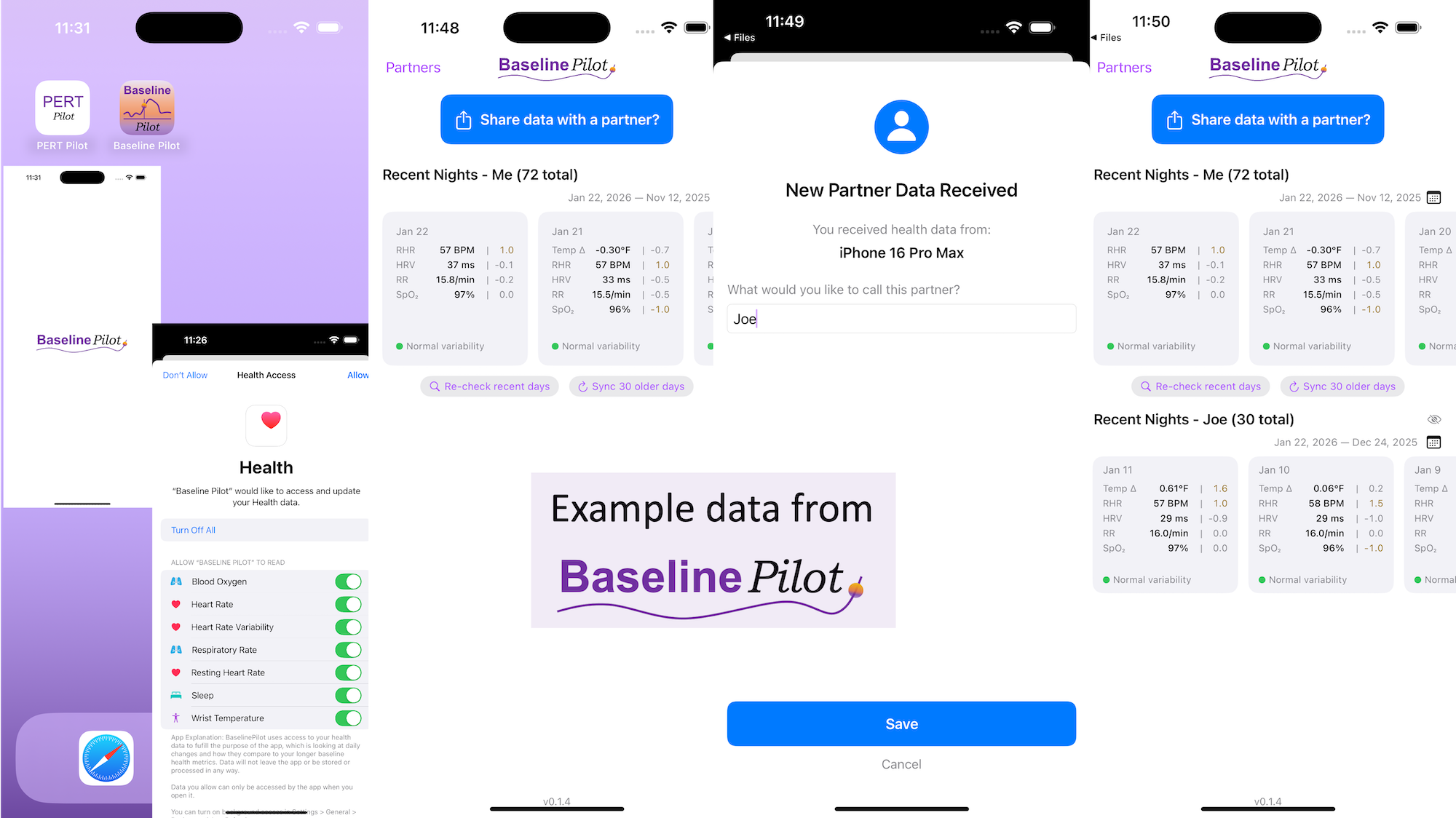
 All of this data is local to the app and not being shared via a server or anywhere else. It makes it quick and easy to see this data and easier to spot changes from your normal, for whatever normal is for you. It makes it easy to share with a designated person who you might be interacting with regularly in person or living with, to make it easier to facilitate interventions as needed with major deviations. In general, I am a big fan of being able to see my data and the deviations from baseline for all kinds of reasons. It helps me understand my recovery status from big endurance activities and see when I’ve returned to baseline from that, too. Plus the spotting of infections earlier and preventing spread, so fewer people get sick during infection season. There’s all kinds of reasons someone might use this, either to quickly see their own data (the Vitals access problem) or being able to share it with someone else, and I love how it’s becoming easier and easier to whip up custom software to solve these data access or display ‘problems’ rather than just stewing about how the standard design is blocking us from solving these issues!
All of this data is local to the app and not being shared via a server or anywhere else. It makes it quick and easy to see this data and easier to spot changes from your normal, for whatever normal is for you. It makes it easy to share with a designated person who you might be interacting with regularly in person or living with, to make it easier to facilitate interventions as needed with major deviations. In general, I am a big fan of being able to see my data and the deviations from baseline for all kinds of reasons. It helps me understand my recovery status from big endurance activities and see when I’ve returned to baseline from that, too. Plus the spotting of infections earlier and preventing spread, so fewer people get sick during infection season. There’s all kinds of reasons someone might use this, either to quickly see their own data (the Vitals access problem) or being able to share it with someone else, and I love how it’s becoming easier and easier to whip up custom software to solve these data access or display ‘problems’ rather than just stewing about how the standard design is blocking us from solving these issues! These are actionable, doable, practical things we can all be doing, today, and not just gnashing our teeth. The sooner we course correct with improved data availability, the better off we’ll all be in the future, whether that’s tomorrow with better clinical care or in years with AI-facilitated diagnoses, treatments, and cures.
These are actionable, doable, practical things we can all be doing, today, and not just gnashing our teeth. The sooner we course correct with improved data availability, the better off we’ll all be in the future, whether that’s tomorrow with better clinical care or in years with AI-facilitated diagnoses, treatments, and cures.  (Thank you).
(Thank you). TL;DR: Instead of arbitrarily lowering or increasing fat or fiber in the diet, measure and estimate what you are consuming first. If you have EPI,
TL;DR: Instead of arbitrarily lowering or increasing fat or fiber in the diet, measure and estimate what you are consuming first. If you have EPI,  If you can shift your mindset from fear and avoidance to curiosity and experimentation, you might discover new skills, solve problems you once thought were impossible, and open up entirely new opportunities.
If you can shift your mindset from fear and avoidance to curiosity and experimentation, you might discover new skills, solve problems you once thought were impossible, and open up entirely new opportunities.
 TLDR: Instead of asking “Which model is best?”, a better question might be:
TLDR: Instead of asking “Which model is best?”, a better question might be:
Recent Comments